Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England
A novel SARS-CoV-2 variant, VOC 202012/01 (lineage B.1.1.7), emerged in southeast England in November 2020 and is rapidly spreading toward fixation. Using a variety of statistical and dynamic modelling approaches, we estimate that this variant has a 43–90% (range of 95% credible intervals 38–130%) higher reproduction number than preexisting variants. A fitted two-strain dynamic transmission model shows that VOC 202012/01 will lead to large resurgences of COVID-19 cases. Without stringent control measures, including limited closure of educational institutions and a greatly accelerated vaccine roll-out, COVID-19 hospitalisations and deaths across England in 2021 will exceed those in 2020. Concerningly, VOC 202012/01 has spread globally and exhibits a similar transmission increase (59–74%) in Denmark, Switzerland, and the United States.
Read the paper in Science here.
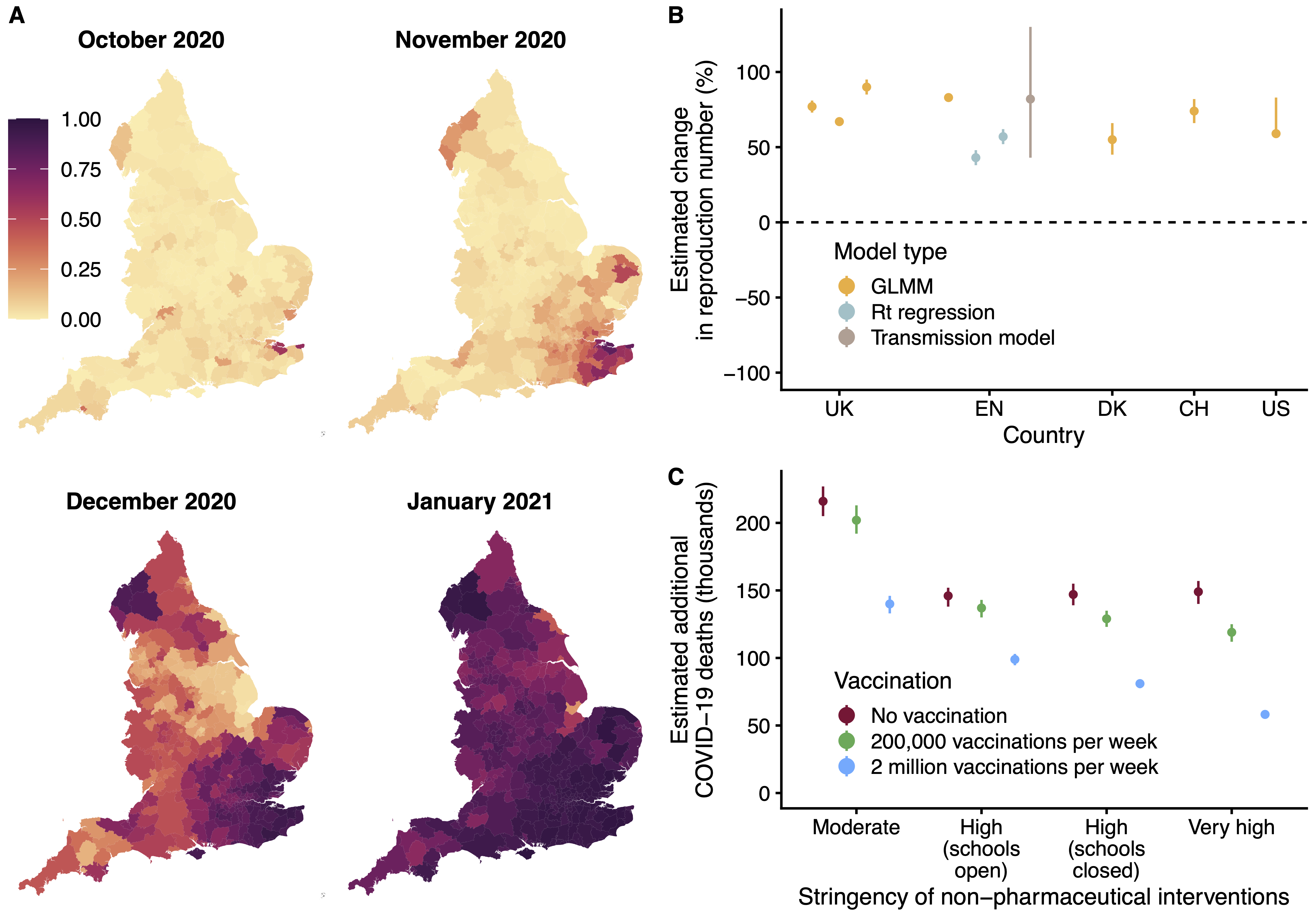
Impact of SARS-CoV-2 Variant of Concern 202012/01. (A) Spread of VOC 202012/01 in England. (B) The estimated relative transmissibility of VOC 202012/01 is similar across the United Kingdom / England, Denmark, Switzerland, and the United States. (C) Projected COVID-19 deaths in England, 15 December 2020–30 June 2021. Vaccine rollout and control measures help mitigate the burden of VOC 202012/01.
Other links